Variation in the karyotype length and genome size of fungus-farming ants
In the recent article “Intraspecific variation in the karyotype length and genome size of fungus-farming ants (genus Mycetophylax), with remarks on procedures for the estimation of genome size in the Formicidae by flow cytometry” published in PLOS ONE, Mariana Neves Moura, Danon Clemes Cardoso, Brenda Carla Lima Baldez, and Maykon Passos Cristiano study the karyotype length and genome size of different populations of two fungus-farming ants Mycetophylax conformis (Mayr, 1884) and Mycetophylax morschi (Emery, 1888). They reveal that the chromosome number and morphology did not vary among the populations of M. conformis or the cytotypes of M. morschi, but karyotype length and genome size were distinct among the populations of these ants. Here, Maykon Passos Cristiano and Danon Clemes Cardoso highlight the main points of their study.
An Interview compiled by Patrick Krapf, Lina Pedraza, and Stephanie Wendt

MNB: Could you tell us a bit about yourself?
DCC and MPC: We are professors at the Department of Biodiversity, Evolution and Environment, at the Federal University of Ouro Preto. Ouro Preto is a World Heritage city in Brazil which was founded at the end of the 17th century. Currently, we are leading the Laboratory of Evolutionary and Population Genetics.
In our lab, we study the evolutionary dynamics that have shaped and still act on biodiversity, in a constantly changing world. Our projects focus on the study of ant genetics and evolution, focusing on the evolutionary history of fungus-farming ants. This group of ants have established a symbiotic relationship with basidiomycete fungi and were presumably originated in non-forest habitats in South America.
Based on sequencing data and insights from different cytogenetic methods, we aim to understand the phylogenetic relationships and the evolution of the genome of these ants. We investigate the impact of past natural selection events and the mechanisms of speciation.
We are currently interested in understanding the impact of eusociality and colonial behavior on the evolution of different morphological, behavioral, and genome traits of the fungus-farming ants.
MNB: Could you briefly outline your research on “Intraspecific variation in the karyotype length and genome size of fungus-farming ants (genus Mycetophylax), with remarks on procedures for the estimation of genome size in the Formicidae by flow cytometry” you published in PlosOne in layman’s terms?
DCC and MPC: Ants are a very diverse insect group and similarly show an astonishing variety of karyotypes. Karyotypes basically define the number of chromosomes and their structure (form) that ultimately underlies their function. Based on our primary aim, looking for the patterns of chromosomal evolution in ants and the process shaping these, we evaluate whether the variation of the karyotype length (sum of the physical measurements of all chromosomes) was positively correlated with DNA amount in the nucleus.


MNB: And what is the take-home message of your work?
DCC and MPC: In this work, we demonstrated that the variation in the DNA amount positively correlates with the size of the chromosomes. This is the second example of such a correlation in fungus-farming ants, and the first example was published by us in 2018 (Cardoso et al. 2018), among populations of Mycetomoellerius holmgreni. This means that changes in the amount of DNA are accumulated across the chromosomes and may occur previous to any changes in chromosome number. Among Formicidae, none study has previously coupled a karyomorphometric approach with genome size data, but such studies are quite common with plants, for instance.
MNB: What was your motivation for this study?
DCC and MPC: Our principal motivation started when we observed that distinct populations of fungus-farming ants showed a variation on the mean karyotype length and this was not at random. Thus, to validate this observation, we started to pursue the genome size estimated for these colonies in order to have an independent and complementary quantification to evaluate the differences observed across colonies.
MNB: What was the biggest obstacle you had to overcome in this project?
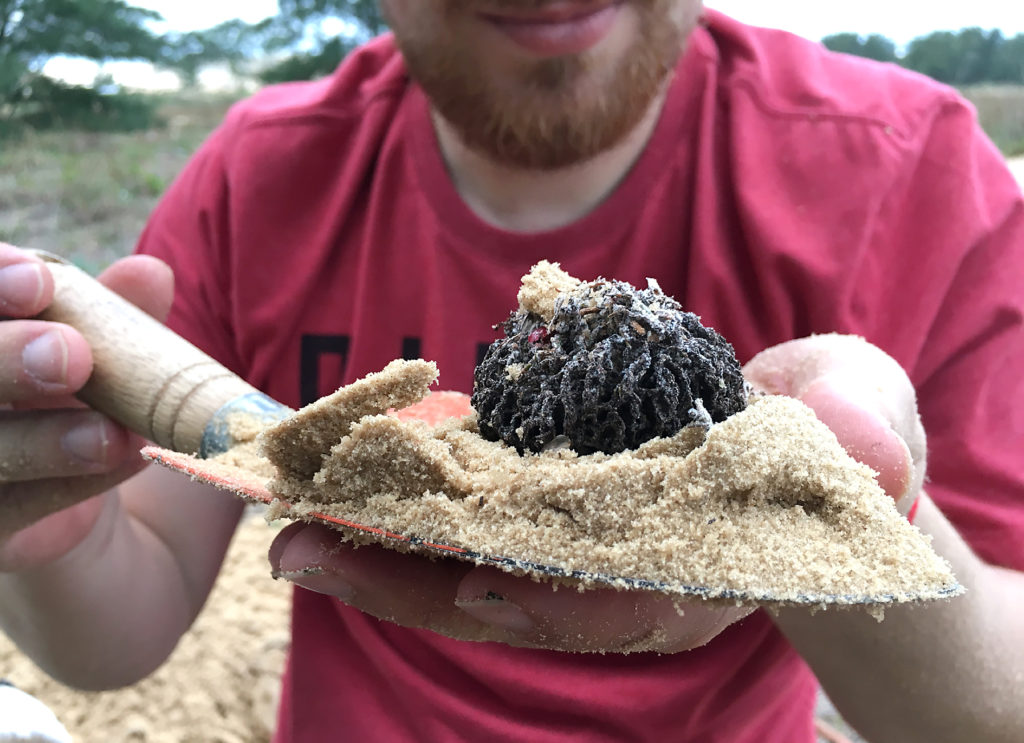
DCC and MPC: The first obstacle to overcome was to establish a protocol for flow cytometry analysis (FCM) at our lab. We started using the previous protocol we have followed in other studies developed in the lab of our partners, but it didn’t work promptly. Because of that, we decided to discuss the protocols of FCM methods applied to studied genome size in ants and included it in our manuscript. We found that distinct methods have been applied in the available studies, so we decided to check, particularly for ants, the three main components of FCM analysis: source tissue, the lysis buffer, and the internal standard. The internal standard is used as a reference to estimate the genome size of the target species.


MNB: Do you have any tips for others who are interested in doing related research?
DCC and MPC: Chromosomes are our passion, but you will need the patience to get good metaphase preparations and all be willing to get fresh material for both cytogenetic and FCM analysis and repeat the trials whatever times it needs.
MNB: Where do you see the future for this particular field of ant research?
DCC and MPC: Ant cytogenetics is an open and interesting field of research. Only a small number of ant species has been studied cytogenetically, and the family Formicidae is an intriguing model for the study of chromosome biology. There is a limited number of research groups that are still working on this subject, which has lost research space with the advance of molecular biology. However, we think there are still innumerous open questions on chromosome biology and karyotype evolution. A fundamental question still on the debate is whether chromosomal changes are the cause or the consequence of lineage differentiation. More and more genomic data has improved cytogenetic analysis and certainly will allow the evaluation of processes shaping chromosomal changes and their role in genome evolution.
References
Moura, M.N., Cardoso, D.C., Lima Baldez, B.C. & Cristiano, M. P. 2020. Intraspecific variation in the karyotype length and genome size of fungus-farming ants (genus Mycetophylax), with remarks on procedures for the estimation of genome size in the Formicidae by flow cytometry. PloS one, 15(8), e0237157.
Cardoso, D.C., Heinze, J., Moura, M.N. & Cristiano, M.P. 2018: Chromosomal variation among populations of a fungus-farming ant: implications for karyotype evolution and potential restriction to gene flow. – BMC evolutionary biology 18: 146.




Recent Comments