Old samples tell new stories


In the recent paper “Genetic identification of Formica rufa group species and their putative hybrids in northern Europe” published in Myrmecological News, Pekka Pamilo and Jonna Kulmuni analysed 14403 workers from 1394 nests in 40 populations and studied species differences and hybridization using allozyme variation in wood ants. With the results, they were able to differentiate between species and to indicate potential areas of hybridization. Interestingly, almost all F. polyctena in Finland are admixed and present only in southern Finland. In contrast, in the northernmost populations, hybridization between F. aquilonia and F. lugubris was detected. These results are important to assess genome-wide selection patterns in hybridizing Formica ants and to understand climate change effects on geographic distribution and hybridization. Here, Pekka and Jonna answer questions and share pictures and videos of the work behind the paper.
An Interview compiled by Alice Laciny, Patrick Krapf, and Roberta Gibson

MNB: Could you tell us a bit about yourself?
PP: I have studied evolution and population genetics of social insects since I started my undergraduate work almost 50 years ago. The preliminary topic of my undergraduate project was genetic differentiation among the species of the Formica rufa group ants. That proved to be challenging with the genetic techniques we had at that time and I ended up including other data in the master’s thesis. Encouraged by my supervisors Rainer Rosengren and Ross Crozier, my research shifted to sociogenetics of ants, but I have returned every now and then to the challenges provided by the Formica rufa group species.
JK: I was Pekka’s last PhD student and as my MSc project picked up on studying a weird Formica population that turned out to be a hybrid. After my PhD, I took speciation and hybridization as the focus of my studies and have developed Formica wood ants as a suitable model for current questions in those fields.

MNB: Could you briefly outline the research you published in Myrmecol. News in layman’s terms?
PP: The Formica rufa group ants are dominant species with great ecological impact in the coniferous forests of northern Europe. The group includes several closely related species that can hybridize with each other. The aim of the work was to estimate how different the species are genetically, whether there are indications of hybridization, and what the geographical differences within a single species might tell us about the history of the populations.
JK: In addition, it is great to get this extensive dataset from the late 1980s published because it will be a valuable point of comparison for later genomic studies of hybridization in these ants.
MNB: What is the take-home message of your work?
PP: Genetic differences between the species are small but consistent in the sense that morphologically identified conspecific populations from different geographical areas tend to cluster together. Possible recent hybridization is indicated by a mismatch between morphological and genetic clustering but it is difficult to detect hybridization that may have taken place in the past history and affected populations within a large geographical area.
JK: To add, we found signatures of hybridization between F. polyctena and F. aquilonia in the south and indication of hybridization between F. lugubris and F. aquilonia in the north.


MNB: What was your motivation for this study?
PP: As mentioned above, I started to work with the Formica rufa group species in my undergraduate work. Some results were published at the end of the 1970s, but many questions remained unsolved. Ten years later I was asked to give a talk on population genetics of this group of ants at the IUSSI congress in 1986. Our methods of allozyme studies had improved and I collected new data to be presented at the congress. The preliminary results showed some interesting genetic patterns, so I decided to continue the work and the samples were collected at the end of the 1980s. This is the material in the current paper. While analyzing the data, a new method – DNA microsatellites – became available and offered more powerful analyses. For this reason, the allozyme data were put aside. However, this was a large data set and recent findings of hybridization and the possibility of range shifts resulting from climate change made us think that it can be valuable to look what the data collected over 30 years ago tell us.
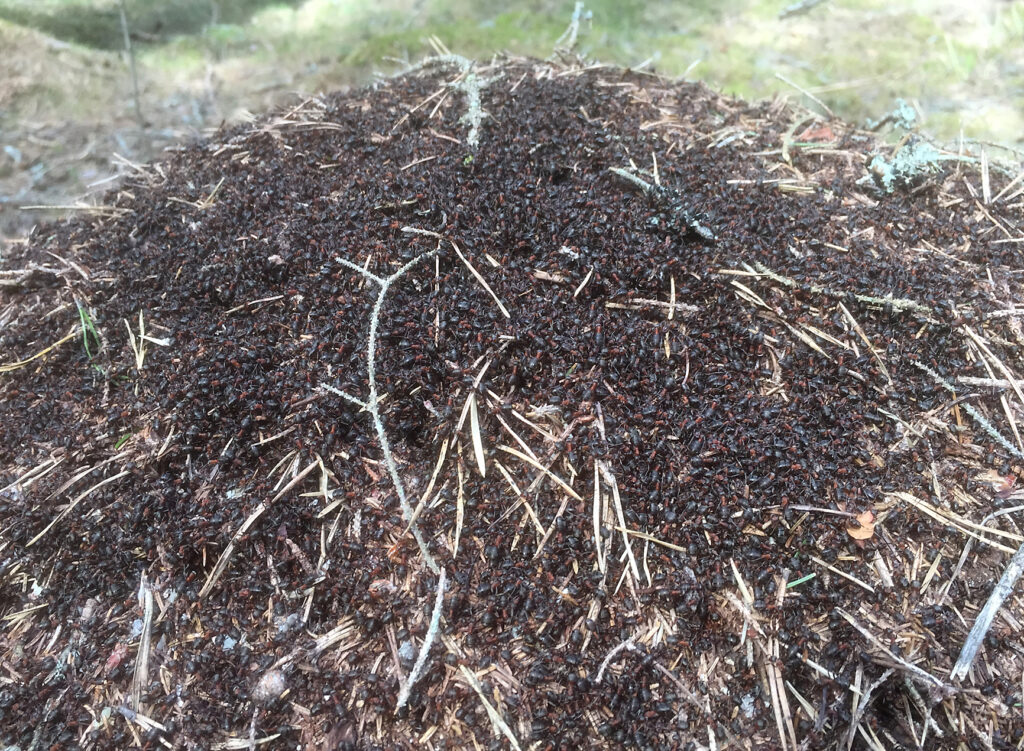
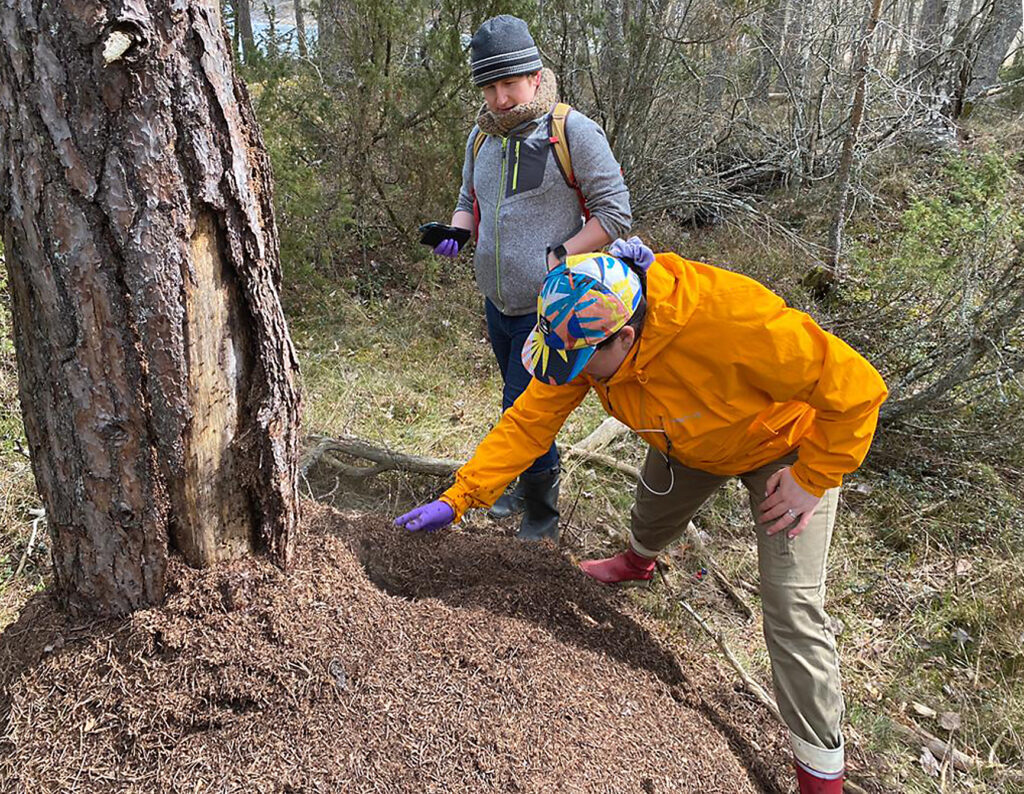
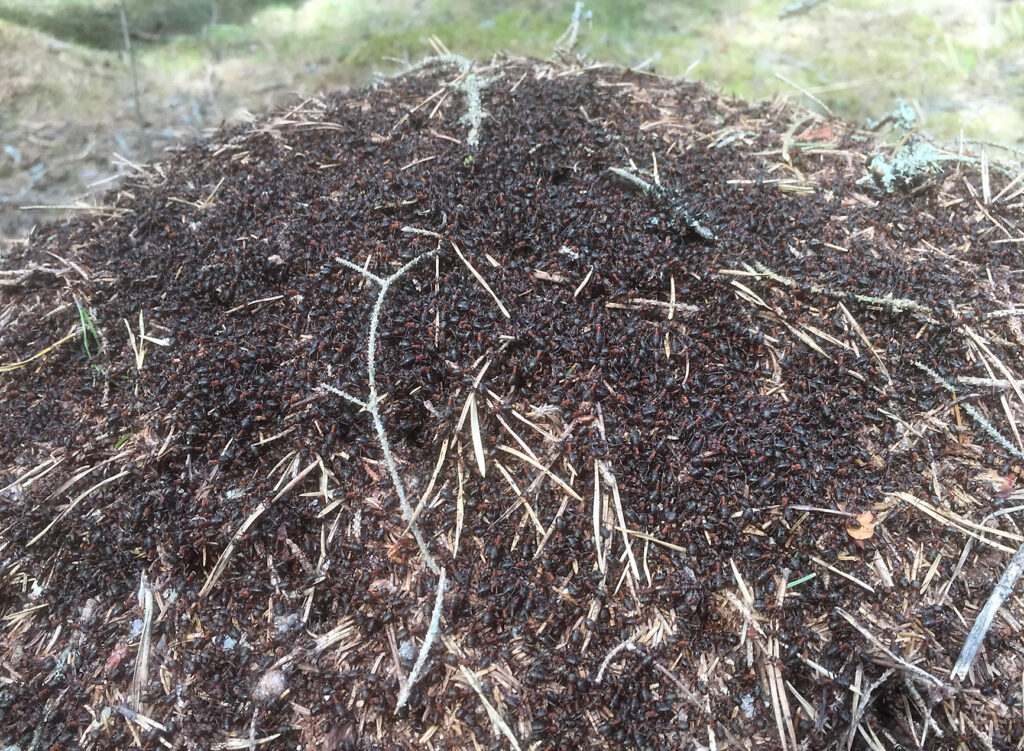
Formica workers on a nest (© Jonna Kulmuni):
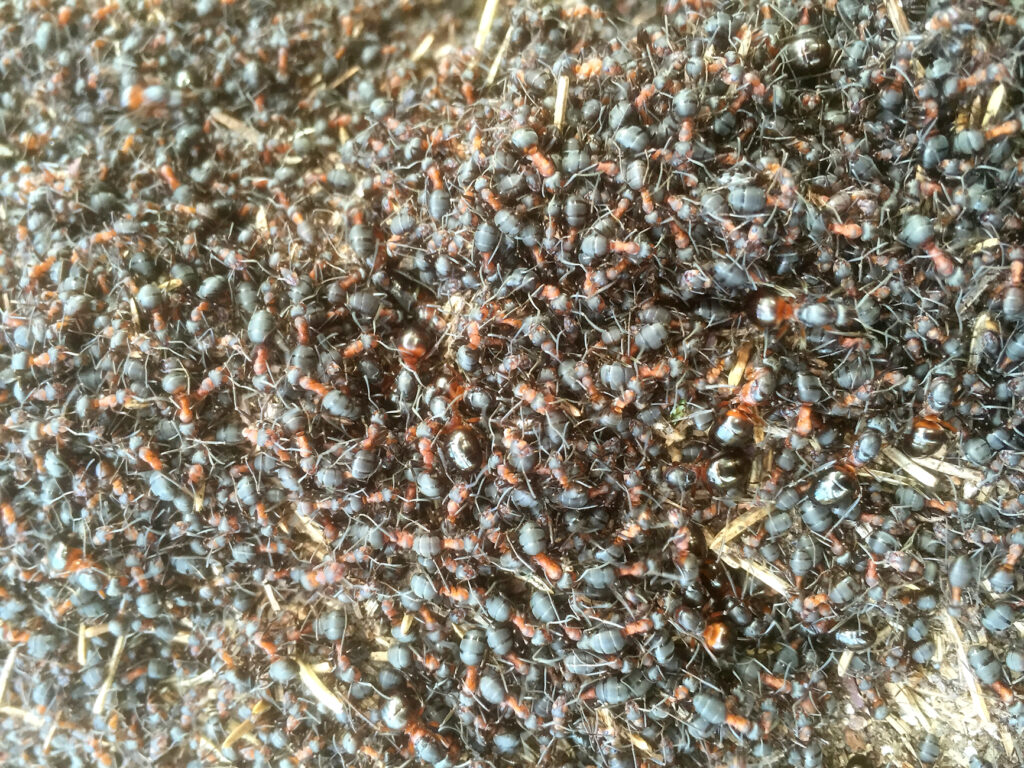
Formica aquilonia – gynes on the nest (© Jonna Kulmuni):
MNB: What was the biggest obstacle you had to overcome in this project?
PP: The power of allozymes to reveal patterns of genetic differentiation is not very strong. This is partly compensated by the size of the data set, but we had to be very careful not to over-interpret the results. Another problem was the morphological identification of the samples. Bernhard Seifert has done important work in describing how to identify species and their hybrids, but our material was collected before his work. Congruence between morphological and genetic results makes our main conclusions reliable but many details remain open.
MNB: Do you have any tips for others who are interested in doing related research?
PP: In order to understand changes that take place in nature we need baseline data as a reference. Similar data sets for studying the geographical distribution of the Formica rufa group ants have been recently collected by national forest inventories (Punttila & Kilpeläinen) and by using citizen science (Jouni Sorvari).
JK: Since genetic data and morphology are two distinct sources of information that don’t always align, both are needed for reliable conclusions about hybridization, especially with a group as difficult as the F. rufa group ants.
MNB: Where do you see the future for this particular field of ant research?
PP: Recent studies on hybridization between F. aquilonia and F. polyctena were the main motivation to analyse the old allozyme data. Both species form large polydomous colonies or supercolonies. We know that supercolonial ants can be fast in invading and spreading into new areas. It is also interesting to see how the populations of such supercolonial ants behave and differentiate in their native ranges and what happens when two such species meet and start hybridizing. There will certainly be more research on hybridization and its consequences using genome-wide data.
JK: In evolutionary biology, ants have been intensively used as models for the evolution of sociality because they represent one of the major transitions. Due to haplodiploidy and social structure, the wood ants are also great models for speciation. Because haploid males reveal natural selection against recessive deleterious allele combinations (i.e. incompatibilities), ants allow mapping of loci underlying reproductive isolation. Moreover, since supercolonial ant populations are mainly closed entities with no or little gene flow from other populations, they allow the study of repeatability of evolution in nature e.g. by studying replicate hybrid populations.







More articles like this, putting old data in new light, should be written. Great article!