Ant phylogenomics

In the article “Ant phylogenomics reveals a natural selection hotspot preceding the origin of complex eusociality”, Romiguier and colleagues* discuss how the evolution of eusociality in ants has allowed ants to become such dominant and successful organisms. The study focuses on the formicoids, a clade of nine subfamilies that exhibit extreme forms of reproductive division of labour, colony size, and worker polymorphism. By sequencing 65 ant genomes, the authors provide a robust phylogeny of the 17 ant subfamilies and show that the controversial leptanillomorph clade is the sister group to all other extant ants. The study also found that the emergence of the formicoids was accompanied by an elevated number of positive selection events. Here, first author Jonathan Romiguier highlights the main points.
* Jonathan Romiguier, Marek L. Borowiec, Arthur Weyna, Quentin Helleu, Etienne Loire, Christine La Mendola, Christian Rabeling, Brian L. Fisher, Philip S. Ward, and Laurent Keller
An Interview compiled and edited by Alice Laciny and Lina Pedraza

MNB: Could you tell us a bit about yourself?
JR: First and foremost, I am an evolutionary biologist interested in how genomes evolve depending on species’ life-history strategies. I always had a particular interest in ants but I started my career in mammal genomics. At this time (2010), mammals were the only animal taxa with several dozens of sequenced genomes for species with very contrasted phenotypes. I consider myself very lucky in having this opportunity and I learned most of what I know in bioinformatics, phylogenomics, and comparative genomics by playing with these mammalian genomes. After having worked on a larger taxonomic scale by estimating genetic diversity determinants across all animals, I started to entirely focus on ants at the University of Lausanne in 2014. I had a great time there and learned a lot. Sequencing technologies started to become more affordable, which opened the doors for sequencing genomes of various ant species. In 2018, I went back to France, to the University of Montpellier as a CNRS researcher, and now focus on my favourite ant species, the harvester ant Messor barbarus.
MNB: Could you briefly outline the research you recently published in layman’s terms?
JR: Ants fascinate too many people because of their high level of sociality. While ant workers are mostly sterile and only live a few years, some ant queens can live several decades and are hyper-fertile, producing colonies of several millions of individuals. Ant genomics first focused on species with a high level of eusociality. The problem is that most of these species belong to a single clade named Formicoids. Without more non-Formicoid genomes, it was difficult to understand the emergence of advanced eusociality, and more generally to resolve the ant tree of life. We filled this gap by sequencing the genome of 65 ant species along the 17 ant subfamilies. This allowed us to clarify the relationships among all ant subfamilies and revealed a natural selection hotspot that occurred before the emergence of Formicoids.


MNB: What is the take-home message of your work?
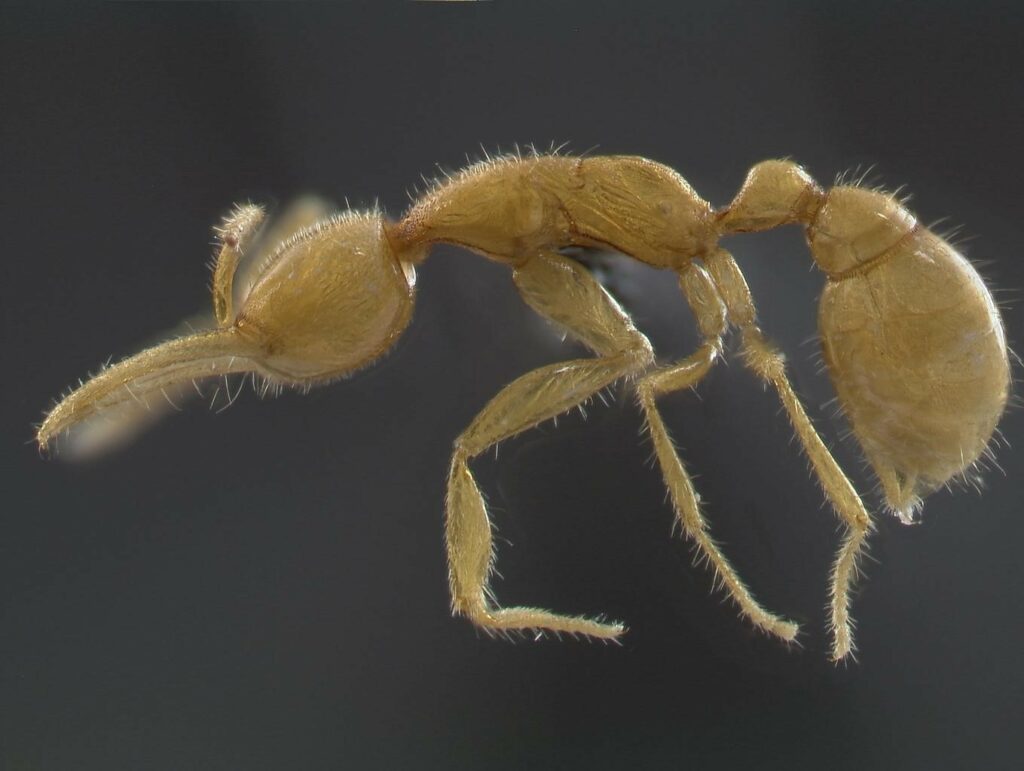
JR: Our phylogenetic analyses support the existence of three main ant groups: Formicoids, Poneroids and Leptanillomorphs. Leptanillomorphs are pale, small, and subterranean ants that represent less than 1% of ant species but form the sister clade of all other ants. Interestingly, the Formicoid emergence was accompanied by an extraordinary natural selection hotspot, including gene functions that are likely to be implicated in key features of eusociality, such as caste differentiation, worker sterility or longevity. These suggest that the molecular foundations of complex eusociality may have rapidly evolved in less than 20 Ma during the Cretaceous.
MNB: What was your motivation for this study?
JR: The rooting of the ant phylogeny has been highly controversial since the advent of molecular phylogenetics, with many different results from well-conducted studies for such an important node. Trying to clarify this conundrum with whole genome sequencing was a motivating challenge, particularly given that it allowed us to collect and observe so many diverse ant groups, which is always a thrill for an ant enthusiast.
MNB: What was the biggest obstacle you had to overcome in this project?
JR: To conduct this study, collecting individuals of some very rare species was mandatory. I could not have achieved this without the help of the great myrmecologist community, particularly my co-authors. For some rare specimens, very low DNA quantities were available, and I cannot thank our lab engineer Christine La Mendola enough for her help.


MNB: Do you have any tips for others who are interested in doing related research?
JR: The best tip is to be prepared to be patient: such a project takes much time. Collecting a large diversity of species means it will not be done in a single field trip and will involve many collaborators. The large amount of data generated by 65 whole sequenced genomes also means a lot of time: every single step, test, or parameter change during the analyses takes a discouraging amount of time, sometimes several months for a minor detail. I started this project in 2015 and published it in 2022. Perseverance and patience are more than useful in a 7-year project.
MNB: Where do you see the future for this particular field of ant research?
JR: These kinds of large comparative genomic studies will become more and more collaborative. The Global Ant Genomics Alliance (GAGA) is the future of ant genomics. It will produce even more ant genomes that will allow us to understand the evolutionary history of ants and the emergence of eusociality in greater detail.






Thank you for reviewing the three articles! Yours, Marc.
See https://blog.myrmecologicalnews.org/2022/09/21/the-first-high-resolution-global-map-of-ant-biodiversity-and-estimates-of-where-future-sampling-could-yield-undiscovered-species/
See https://blog.myrmecologicalnews.org/2023/03/30/ant-abundance-biomass-and-distribution/