Small specimens provide big data – using Pheidole ants for museomics
In the recently published article “Revisiting museum collections in the genomic era: potential of MIG-seq for retrieving information from aged minute dry specimens of ants (Hymenoptera: Formicidae) and other small organisms” published in Myrmecological News, Katsuyuki Eguchi and colleagues tested the applicability of a high-throughput sequencing-based method on museomics (genomic data derived by the use of organisms stored in natural history collections). Here, Hannah Weigand highlights their main points.
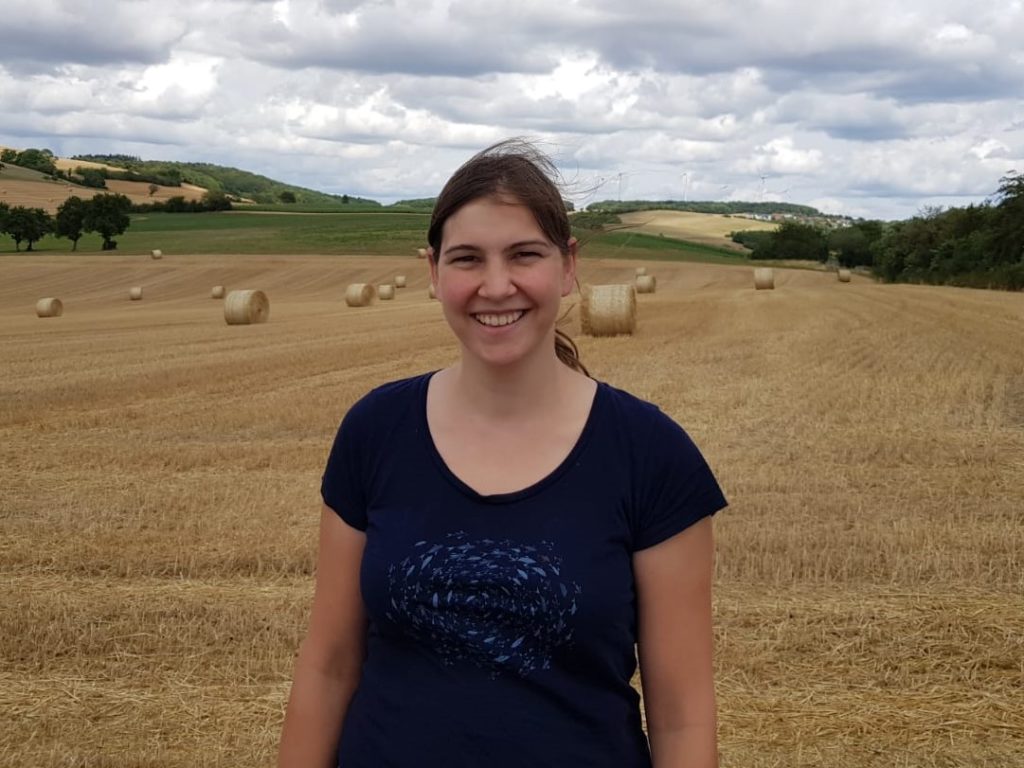
A View by Hannah Weigand

In the last years, new high-throughput sequencing (HTS) platforms and applications have opened the fields of phylogenomics and population genomics for samples from museum collections (i.e. museomics). However, long-term storage conditions in natural history collections often are not adequate to preserve specimens for genetic studies. As a result, low DNA quality and quantity can form major challenges for HTS-based applications. These challenges can be even more pronounced for small target organisms, such as ants.
So-called reduced representation HTS-based methods can be applied to deal with non-optimal storage conditions. Those methods do not need a completely sequenced genome but rely on homologous proportions of the genome which are sequenced. In principle, this means that lower DNA quantities and more fragmented DNA can contribute to successful molecular data collection. As the homologous DNA fragments are (ideally) shared among (all) analysed specimens, the reduced representation methods vary mainly in the way how a specific fraction of the genome is selected (or reduced).
In the article published in Myrmecological News by Eguchi et al. (2020), the reduced representation method MIG-seq (multiplexed ISSR genotyping-by-sequencing) was tested for its applicability in museomics. In this approach, simple sequence repeats (SSR; i.e. microsatellites) were used as recognition motives for sequencing, while the genetic variance (e.g. SNPs, i.e. single nucleotide polymorphisms) placed between two microsatellites (I; inter) was analysed. The number of analysed homologous loci can be controlled by specifying both microsatellite motives (e.g. (ACT)4TG) and the size range spacing both microsatellite motives (e.g. >250 bp). In most cases, homologous sequences are expected to be framed by the same microsatellite motives (though with different numbers of repeats) and separated by an approximately equal distance; thus, the method allows to find the same loci for different specimens.
While homologous loci are usually obtained within a species relatively easily, mutations accumulating with deeper divergence time will result in a lower overlap between species. Additionally, the quality or quantity of the DNA from museum specimens might be insufficient for the method. Hence, Eguchi et al. tested the applicability of MIG-seq for museomics by analysing the phylogenetic relationships of 55 specimens from 46 species of the ant genus Pheidole. Most samples were retrieved from dry-mounted specimens with 10 to 23 years of age, fewer specimens were conserved in ethanol (75% or absolute). A total of approx. 50 million sequences was produced with the Illumina MiSeq technology, resulting in between approx. 260,000 and 3,700,000 reads per sample. Different quality-filtering settings were tested for locus clustering and SNP identification, generating a final dataset with 4,849 loci and 36,862 SNPs (648 – 15,778 per sample). While DNA quality and quantity did not correlate with the storage time of samples, longer storage times still resulted in lower numbers of reads and SNPs produced. This might be particularly problematic for phylogenetic or population genetic studies if storage conditions are linked to certain taxonomic groups, for example, if samples from one taxon, clade, or population are stored for a longer period of time than others. Future study designs of MIG-seq applications should take this factor into account. The study of Eguchi et al. could have benefited from a more controlled setup for a better understanding of data quality. For example, double sequencing of selected specimens would have allowed them to calculate genotyping error rates directly.

In a next step, a phylogenetic tree reconstruction was made to evaluate how well MIG-seq data reflect known phylogenetic relationships within the genus Pheidole. Large parts of the Bayesian tree were in agreement with Economo et al. (2015), who based their study on nine loci including mitochondrial and nuclear markers. However, nodes reflecting deep splitting events were not well resolved in the current reconstruction, probably due to a low overlap of homologous loci. This is in line with the high level of missing data reported in Eguchi et al. (2020). The final dataset used a minimal coverage of 10% of the individuals per locus, thus allowing up to 90% of missing data. More stringent settings with reduced missing data (minimal coverages of 50% to 80% tested) resulted in a very low number of retrieved homologous loci (0 to 47) and SNPs (0 to 421) for the final analysis.
In summary, while the method shows to be problematic for phylogenetic studies analysing older splitting events (nodes of 14 Mya were well resolved), and to be potentially biased by long storage times of samples under suboptimal conditions, the MIG-seq approach still seems to be a method which can be well applied in museomics even for small specimens such as ants. After an initial investment cost in sequencing primers, the method is relatively cost-efficient (4 – 26 US dollars per sample, depending on the level of multiplexing) and is less laboratory demanding than comparable approaches (e.g. ddRAD-Seq).
Further reading
Economo, E.P., Sarnat, E.M., Janda, M., Clouse, R., Klimov, P.B., Fischer, G., Blanchard, B.D., Ramirez, L.N., Ander-sen, A.N., Berman, M., Guénard, B., Lucky, A., Rabeling, C., Wilson, E.O. & Knowles, L.L. 2015b: Breaking out of biogeographical modules: range expansion and taxon cycles in the hyperdiverse ant genus Pheidole. – Journal of Biogeography 42: 2289-2301.
Eguchi, K., Oguri, E., Sasaki, T., Matsuo, A., Nguyen, D.D., Jaitrong, W., Yahya, B.E., Chen, Z., Satria, R, Wang, W.Y. & Suyama, Y. 2020: Revisiting museum collections in the genomic era: potential of MIG-seq for retrieving phylogenetic information from aged minute dry specimens of ants (Hymenoptera: Formicidae) and other small organisms. – Myrmecological News 30: 151-159.
Suyama, Y. & Matsuki, Y. 2015: MIG-seq: an effective PCR-based method for genome-wide single-nucleotide polymorphism genotyping using next-generation sequencing platform. – Scientific Reports 5: art. 16963.
Wachi, N., Matsubayashi, K. W., & Maeto, K. 2018: Application of next‐generation sequencing to the study of non‐model insects. – Entomological Science 21: 3-11.




Recent Comments